Maria Tampakaki¹˒²˒³, Eleftheria Tzamali³, Giorgos Tzedakis³, Alejandros Andrianopoulos²˒⁴, Eirini Makrygiannaki², Vangelis Sakkalis³, Joseph Papamatheakis⁵˒⁶, Giannis Zacharakis²
¹ University of Crete, School of Medicine, Heraklion Crete, Greece
² Foundation For Research and Technology-Hellas, Institute of Electronic Structure and Laser, Heraklion Crete, Greece
³ Foundation For Research and Technology-Hellas, Institute of Computer Science, Heraklion Crete, Greece
⁴ University of Crete, Department of Physics, Heraklion Crete, Greece
⁵ University of Crete, Department of Biology, Heraklion Crete, Greece
⁶ Foundation For Research and Technology-Hellas, Institute of Molecular Biology and Biotechnology, Heraklion Crete, Greece
Introduction
Glioblastoma (GBM) progression emerges from complex, multidimensional interactions between heterogeneous tumor cells and their microenvironment, motivating the development of robust preclinical models. While in vivo systems capture this complexity, they limit experimental control, whereas in vitro models enable controlled investigation of avascular tumor dynamics¹˒². Integrating multiscale imaging data with mathematical modelling offers a path to systematically link in vitro observations to environment-specific in vivo growth behavior.
Methods
The U87MG GBM cell line stably expressing luciferase was used to generate 3D spheroids in ULA plates and to establish intracranial (IC) and subcutaneous (SC) xenografts in immunodeficient NSG mice (5–6 months old). In vitro spheroid growth was monitored using wide-field microscopy and in vivo tumor progression was followed longitudinally using the φ-eye™ optical imaging system (BIOEMTECH). Bioluminescence imaging was acquired after intraperitoneal administration of D-luciferin (150 mg/kg) at standardized time windows. Tumor and spheroid areas were quantified using Fiji. Growth dynamics were analyzed using linear, Gompertz, and logistic models. In addition, mathematical methods were formulated to capture key biological features underlying growth differences across experimental conditions.
Results/Discussion
Longitudinal imaging revealed distinct growth regimes across experimental environments. In vitro GBM spheroids showed approximately linear area expansion, consistent with avascular growth under diffusion-limited conditions and contact inhibition³˒⁴. IC tumors exhibited rapid early expansion followed by saturation, resembling logistic growth likely driven by spatial constraints and brain-specific microenvironmental factors. In contrast, SC tumors displayed a pronounced lag phase of 20–30 days before transitioning to rapid expansion, indicating delayed adaptation to the host tissue (Fig1). Excised tumors exhibited extensive edema and increased vascularization. Phenomenological growth models captured environment-dependent dynamics and indicated doubling times varying from one to a few days across conditions (Fig1). These findings motivate biophysical mathematical models that explicitly account for microenvironmental effects to enable predictive in vitro–in vivo extrapolation.
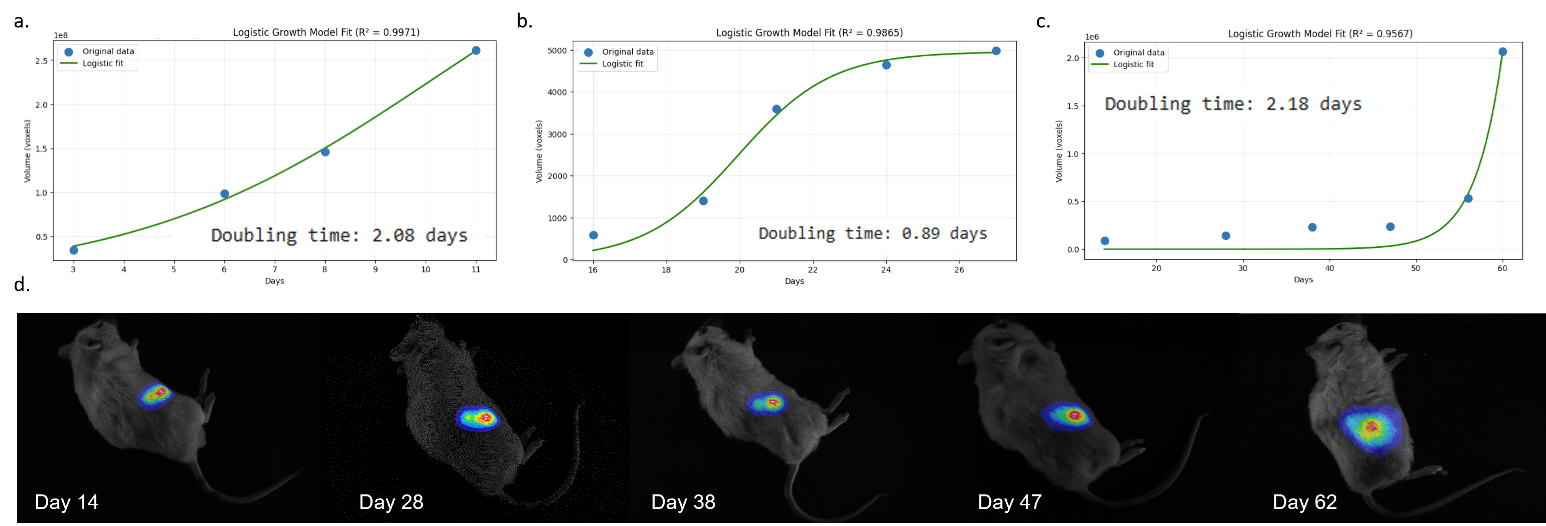
Figure 1: Spatiotemporal growth of U87MG cells in vitro and in vivo. Growth dynamics of U87MG cells in 3D spheroids in vitro (a), intracranial xenografts (b), and subcutaneous xenografts (c). Panel (d) shows longitudinal bioluminescence imaging (BLI) of subcutaneous tumor growth, illustrating the temporal evolution of tumor burden.
Conclusion
This study presents a longitudinal imaging framework to compare glioblastoma growth across in vitro, subcutaneous, and intracranial models. By identifying distinct growth regimes linked to microenvironmental context, the work establishes a validation pipeline for future integration of advanced in vitro systems, perturbation studies, and mathematical modelling toward predictive in vitro–in vivo extrapolation.
Novelty
Longitudinal imaging reveals distinct, environment-dependent GBM growth regimes across in vitro, SC, and IC environments within a unified framework.
Impact
Provides a translational imaging pipeline to support biophysical modelling–driven in vitro–in vivo mapping and predictive GBM research.
Disclosure
Neither I nor any of my co-authors have any financial interests or relationships to disclose in relation to the subject of this presentation
Acknowledgment
The authors would like to thank BIOEMTECH for providing access to the in vivo optical imaging system φ-eye™. This work was supported by the H2020 FETOPEN project Dynamic (GA-863203), the HORIZON Europe EIC Pathfinder Open projects Codaflight (GA-01047263) and SWOPT (GA-101046667) and the HORIZON Europe EDIH project SmartHealth (GA-101083630).
References
[1] Hum NR, Sebastian A, Gilmore SF, He W, Martin KA, Hinckley A, Dubbin KR, Moya ML, Wheeler EK, Coleman MA, Loots GG. Comparative Molecular Analysis of Cancer Behavior Cultured In Vitro, In Vivo, and Ex Vivo. Cancers (Basel). 2020 Mar 14;12(3):690. doi: 10.3390/cancers12030690; PMID: 32183351; PMCID: PMC7140030.
[2] K. Camphausen, B. Purow, M. Sproull, T. Scott, T. Ozawa, D.F. Deen, & P.J. Tofilon, Influence of in vivo growth on human glioma cell line gene expression: Convergent profiles under orthotopic conditions, Proc. Natl. Acad. Sci. U.S.A. 102 (23) 8287-8292, ttps://doi.org/10.1073/pnas.0502887102 (2005).
[3] Oraiopoulou ME, Tzamali E, Tzedakis G, Vakis A, Papamatheakis J, Sakkalis V. In Vitro/In Silico Study on the Role of Doubling Time Heterogeneity among Primary Glioblastoma Cell Lines. Biomed Res Int. 2017;2017:8569328. doi: 10.1155/2017/8569328. Epub 2017 Oct 3 PMID: 29226151; PMCID: PMC5684616.
[4] Jagiella N, Müller B, Müller M, Vignon-Clementel IE, Drasdo D (2016) Inferring Growth Control Mechanisms in Growing Multi-cellular Spheroids of NSCLC Cells from Spatial-Temporal Image Data. PLOS Computational Biology 12(2): e1004412. https://doi.org/10.1371/journal.pcbi.1004412
